Explorer provides a single integrated framework where researchers can explore a wide range of diverse yet interconnected HD scientific data. CHDI and its partners curated and analyzed data from hundreds of HD studies spanning complementary experimental data types using standard, controlled vocabularies and consistent methodologies. Data sources include published literature and ‘omics studies, as well as dozens of previously unreleased CHDI in-life pharmacological studies performed by CRL and Psychogenics.
Data provenance is universally maintained, and experimental meta-data are mapped to centralized HD-specific catalogs of shared elements (e.g. HD Mouse Model Catalog and HD Gene Catalogs). The rich interconnections between data sources, experimental data and reference catalogs allow users to pivot and identify a spectrum of both primary sources and experimental data related to their current inquiry. While Explorer is designed as an interactive tool, complete copies of component datasets are available on the HDinHD Downloads tab for labs who wish to incorporate batch data to facilitate internal data mining efforts.
HD Explorer, enables researchers to browse and search directly for relevant and related primary studies as they look to form, explore and refine scientific hypotheses. There are a number of entry points.
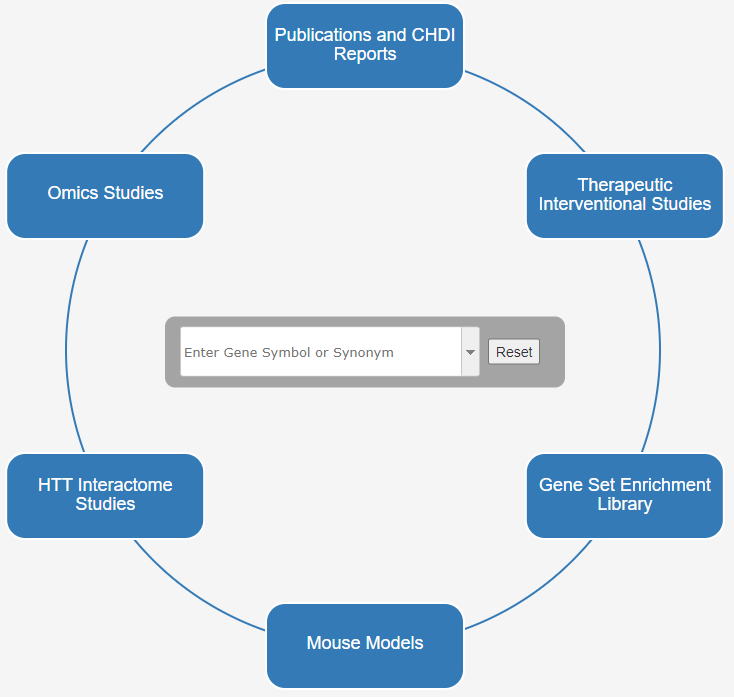
Each box on the Explorer entry circle, as well as the central search box, provide distinct portals into HD Explorer.
- Publications and Reports provides access to >1600 HD-related PubMed articles and CHDI Reports. Reports are bi-directionally linked to experimental data detailed across HD Explorer.
- The Omics Study portal provides access to >250 HD-related omics studies identified in GEO, Array Express and PRIDE. Curated experimental data is presented and links are provided to individual gene sets resulting from analysis of each study
- The Therapeutic Interventions portal provides access to >375 PubMed articles and 66 previously unreleased CHDI reports describing in-life studies (largely with behavioral) readouts. Curated experimental data – including treatment category, treatment name, HD model system – are presented, and each experiment links to a primary source document, many of which are available for download. The gene targets of these intervention studies are linked to the HD Gene Catalog, and HD mouse models are linked to the HD Mouse Model Catalog.
- The Htt Interactome portal contains curation results for 225 PubMed articles detailing >9000 WT and mHTT protein-protein interaction reports. Both positive and negative interactions are presented. Interactor proteins are linked to the HD Gene Catalog.
- The Gene Set Enrichment Library (HDSigDB) contains >2300 gene sets generated from the curation and analysis of HD-related studies. The core of HDSigDB was derived from the curation and analysis of HD and triplet-repeat expansion disease studies deposited in GEO and ArrayExpress. Additional sources of gene sets include: PRIDE, selected PubMed articles and selected pathways. Each gene set links back to its source, and a link is provided to download individual gene sets as lists of both human and mouse gene symbols.
- The HD Mouse Catalog is comprised of >130 distinct HD mouse models described in the literature. Individual mouse models are bi-directionally linked to HD experimental data detailed within HD Explorer that utilize that mouse model.
- Gene-specific pages within the HD Gene Catalog bi-directionally link to complementary experimental data within Explorer and to visualization and reporting tools within HDinHD.
In the coming months, HD Explorer will be extended to include complementary experimental data types, including the HD perturbation literature and previously unreleased internal CHDI HTT Quantification studies from satellite cohorts of the CHDI Therapeutic Intervention studies described above.
In addition to inclusion of new data types, CHDI plans to incrementally update existing Explorer datasets as new research is published.
Access the HD Explorer:


