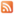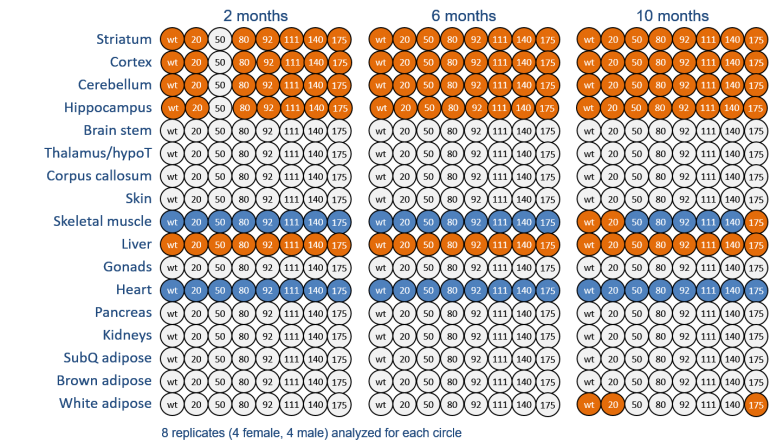
In order to gain a deeper understanding of the pathophysiological mechanisms in Huntington’s disease (HD), we compared the proteomes between wild-type and heterozygous Huntingtin knock-in mice with increasing CAG repeat lengths in a number of different brain regions and peripheral tissues at three different ages (2, 6, and 10 months). The analysis of more than 1,200 tissue samples with on average 8,000 quantified proteins comprises one of the largest global, quantitative proteomics studies published so far. More importantly, it allows a systematic analysis of pathways and interaction networks on the protein level, the identification of novel target candidates, and provides a comprehensive resource for training of system biology models. Part of the data is described in the paper
- Peter Langfelder, Jeffrey P Cantle, Doxa Chatzopoulou, Nan Wang, Fuying Gao, Ismael Al-Ramahi, Xiao-Hong Lu, Eliana Marisa Ramos, Karla El-Zein, Yining Zhao, Sandeep Deverasetty, Andreas Tebbe, Christoph Schaab, Daniel J Lavery, David Howland, Seung Kwak, Juan Botas, Jeffrey S Aaronson, Jim Rosinski, Giovanni Coppola, Steve Horvath, X William Yang: “Integrated genomics and proteomics define huntingtin CAG length-dependent networks in mice”, Nature Neuroscience 19 (2016) 622-633. PMID: 26900923.

The results of the large-scale proteomics study are now accessible through the novel data repository termed HD Proteome Base. The user-friendly web portal allows the researcher to query for proteins and to visualize their expression across the CAG repeat length series and across different tissues. HD Proteome Base is based on MaxQB, which was developed in collaboration with the Max-Planck Institute of Biochemistry in Martinsried, Germany (Schaab et al., MCP 2012, M111. 014068, PMID: 22301388 ![]() ). Whereas MaxQB serves as generic repository for quantitative proteomics data, HD Proteome Base will focus on HD-related experiments. With this effort, we hope to help multiple research teams across the globe to gain a better understanding of the huntingtin biology and potentially contribute to the development of new therapies for HD.
). Whereas MaxQB serves as generic repository for quantitative proteomics data, HD Proteome Base will focus on HD-related experiments. With this effort, we hope to help multiple research teams across the globe to gain a better understanding of the huntingtin biology and potentially contribute to the development of new therapies for HD.
Access the HD Proteome Base:


